|
My_graph <- create_graph(nodes_df = nodes, edges_df = edges)Ī GraphViz object requires a few more steps to save as an image file, such as a PNG: # export graphĪ more complex diagram, specifying different colors for some nodes, and using subgraph clustering. library(DiagrammeR) library(DiagrammeRsvg) library(rsvg)Ī simple diagram: # define nodes dataframeĮdges <- create_edge_df(from = c(1, 2, 3, 3), You can also use the Export functionality in RStudio to save graphs. You must begin by specifying graph in this function (see example below).įor exporting/printing/saving your diagrams, the package DiagrammeRsvg can be useful. Diagrams specified with mermaid must pass a valid diagram specification to the function mermaid(). The mermaid library is also implemented in DiagrammeR.At the minimum, you will need graph, node, and edge statements (see example below). Diagrams specified with Graphviz must pass a valid diagram specification in DOT to the function grViz(). DOT is also used by Graphviz, which is implemented in DiagrammeR.With render_graph you can also output your graph in DOT (graph description language). These include create_graph for defining the structure of your diagram, and render_graph for printing your diagram in the RStudio viewer. 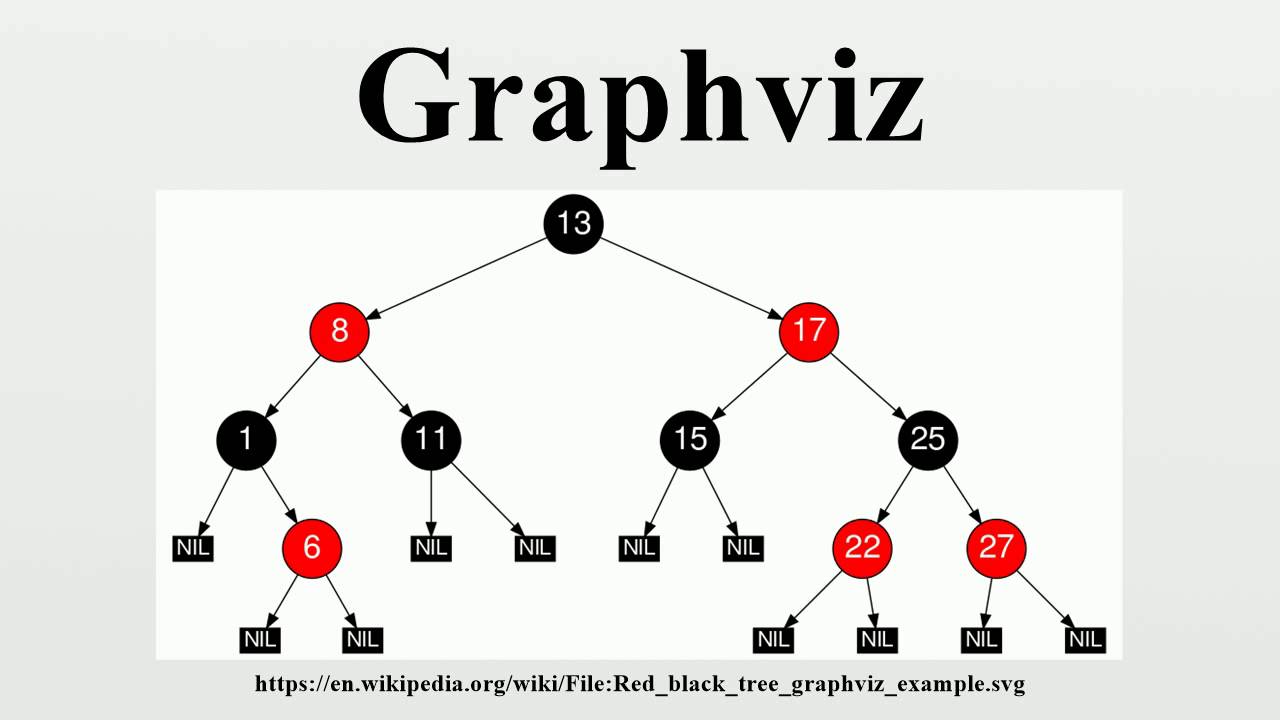
The package contains R functions that help you create a diagram. 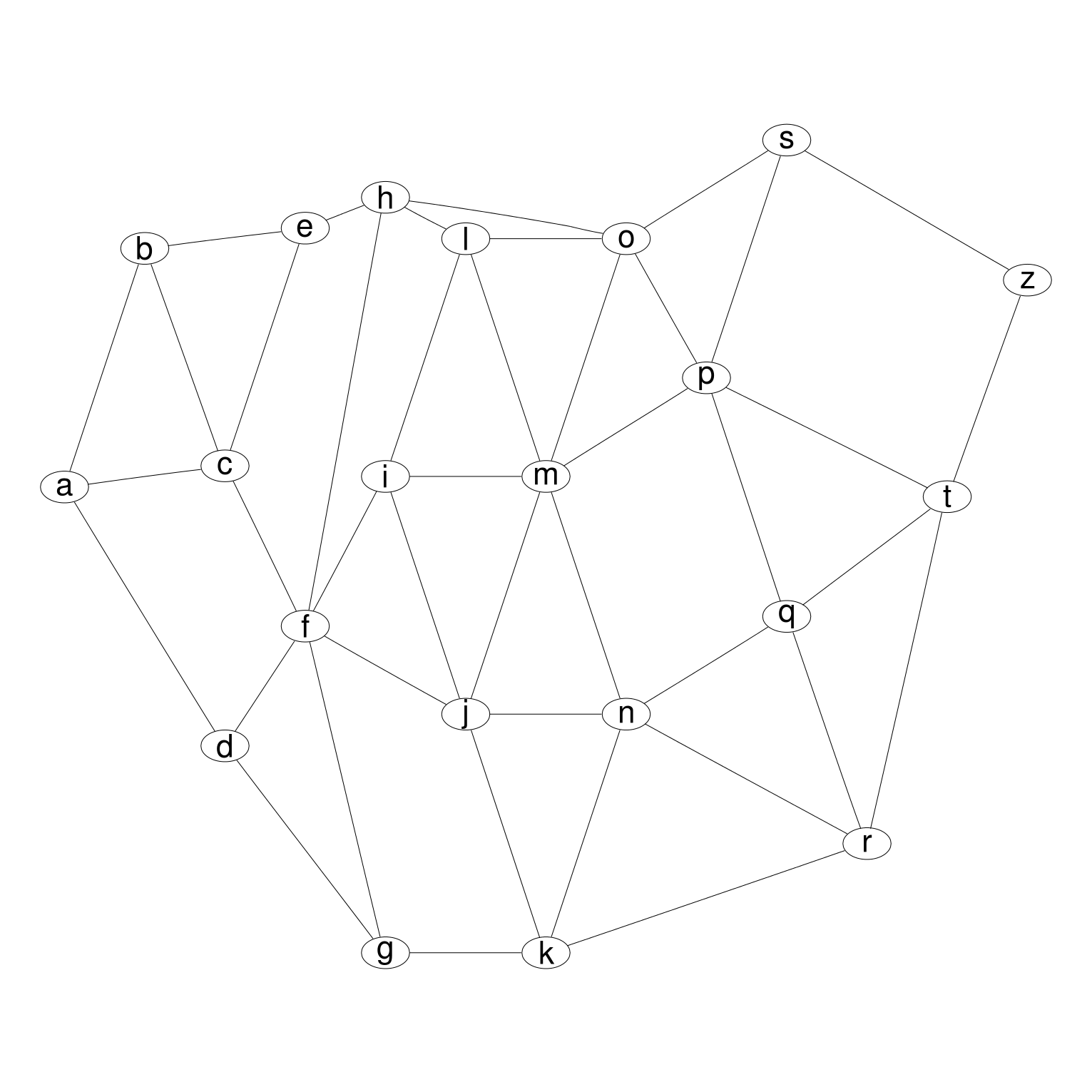
There are three main ways to use the DiagrammeR package: You can check if it is installed with a command like dpkg -s graphivz. The R package DiagrammeR makes it much easier to create high quality figures and diagrams in situations like these. Graphviz is a standard package on many linux distributions. of graph manipulation, notably igraph which also has bindings for R and C++. Package in your R session.Have you ever needed to create a visualization of a research process or statistical model that isn’t directly plotted from data? For example, a conceptual diagram, mind map, flowchart of your research process, or statistical model diagram. Graphviz provides us with more sophisticated layout CAUTION: Do NOT name. Using files of these types in RStudio provides the advantage of syntax coloring and allowing a quick preview of the diagram (via the Preview button, which is activated in RStudio for these types of files). R (>= 2.6.0), methods, utils, graph, gridīiocGraph, BioMVCClass, CellNOptR, flowCL, flowMerge, maEndToEnd, MineICA, netresponse, paircompviz, pathRender, ROntoTools, SplicingGraphs, TDARACNEĪpComplex, biocGraph, BiocOncoTK, bnem, chimeraviz, CytoML, dce, DEGraph, EnrichmentBrowser, flowWorkspace, GeneNetworkBuilder, GOstats, hyperdraw, KEGGgraph, MIGSA, mirIntegrator, mnem, OncoSimulR, ontoProc, paircompviz, pathview, Pigengene, qpgraph, SplicingGraphs, trackViewer, TRONCOĪ4, altcdfenvs, annotate, Category, CNORfeeder, CNORfuzzy, DEGraph, flowCore, geneplotter, GlobalAncova, globaltest, GSEABase, MLP, NCIgraph, NCIgraphData, OmnipathR, pkgDepTools, RBGL, RBioinf, rBiopaxParser, Rtreemix, safe, SNAData, SPIA, SRAdb, Streamer, topGO, ViSEAGO, vtpnet RStudio has integrated file support for Graphviz and mermaid files. To view documentation for the version of this package installedĪ New Interface to Plot Graphs Using Rgraphviz 
Author: Kasper Daniel Hansen cre, aut, Jeff Gentry aut. If (!require("BiocManager", quietly = TRUE))įor older versions of R, please refer to the appropriate Interfaces R with the AT and T graphviz library for plotting R graph objects from the graph package. To install this package, start R (version
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
